Software and data
Over the years I have created or helped to create a number of Software and scripts and novel datasets for environmental analyses at various scales. I usually make all data and code that I have been involved in creating available as openly as possible, for example see my uploads on Zenodo.
Code
ibis.iSDM - Modelling framework for creating Integrated Biodiversity distribution scenarios

The ibis.iSDM package provides a series of convenience functions to fit integrated Species Distribution Models (iSDMs). With integrated models we generally refer to SDMs that incorporate information from different biodiversity datasets, external parameters such as priors or offsets or constrains such as dispersal parameters in future predictions. The ibis.iSDM package has been publicly released and published as Jung 2023.
LecoS - Land cover statistics QGIS plugin

LecoS is a Python plugin for the QGIS GIS software suite. It converts classified raster layers to arrays using the powerful numpy library and - based on a Connected Component Labeling approach - allows to further identify class patches and the calculation of landscape metrics. The use can choose to calculate single or several metrics for the raster classes.
Manuscript | Code | QGIS | Blog post
QSDM - Species Distribution Modelling for the QGIS Processing Toolbox

A while ago I created a python plugin with a series of helper functions to conduct species distribution modelling (e.g. trained single class classifiers) for the QGIS software suite. The plugin is released and functional, but due to a lack of time commitment likely won’t work on latest QGIS versions (> 3.0) anymore. The QSDM package adds the following functionalities to the QGIS Processing toolbox:
Data Analysis
- Calculate Niche Overlap Statistics
- Range Shifts
Data Preperation
- Create Species Richness Grids
- Data Transformation (simple)
- Raster Unification
Species Distribution Modelling
- MAXENT (Parameter Preperation)
- Maximum Entropy Modelling (semi-automatic modelling)
- Maximum Entropy Modelling (Manual Configuration of parameters)icarus - R package for spatial analysis convenience and analysis functions
I created a package to host a number of convenience functions for handling and manipulating spatial datasets in R. Mainly for my personal use, but hopefully the functions will prove useful for others as well
Data
Areas of conservation importance for biodiversity, carbon and water
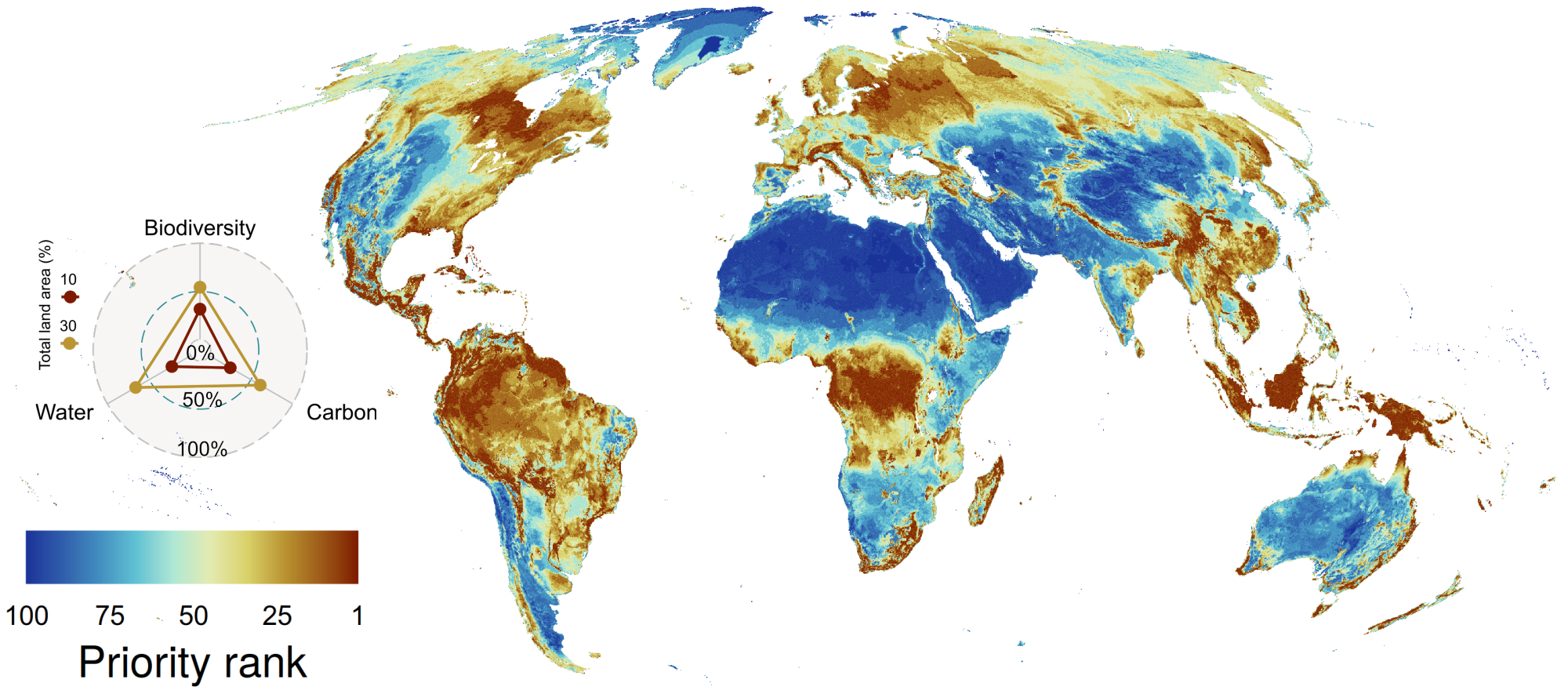
Provided are global conservation importance layers that integrated the best available global biodiversity, carbon and water data at the time. In particular, it includes for the first time a representative set of about half of all plant species globally as well as above, below and soil organic carbon at risk from land use change. Different combinations of layers integrating biodiversity, carbon and/or water value are presented, based on the protected area network at the time (covering ~16% of terrestrial land area) or starting from a blank state. The maps highlight the upper potential value of any global terrestrial land area for conserving these valuable assets and presented as ranking from most (1) to least importance (100).
Interactive Explorer | GEE Assets | Code | Data | Manuscript
Global terrestrial and marine habitat mapping
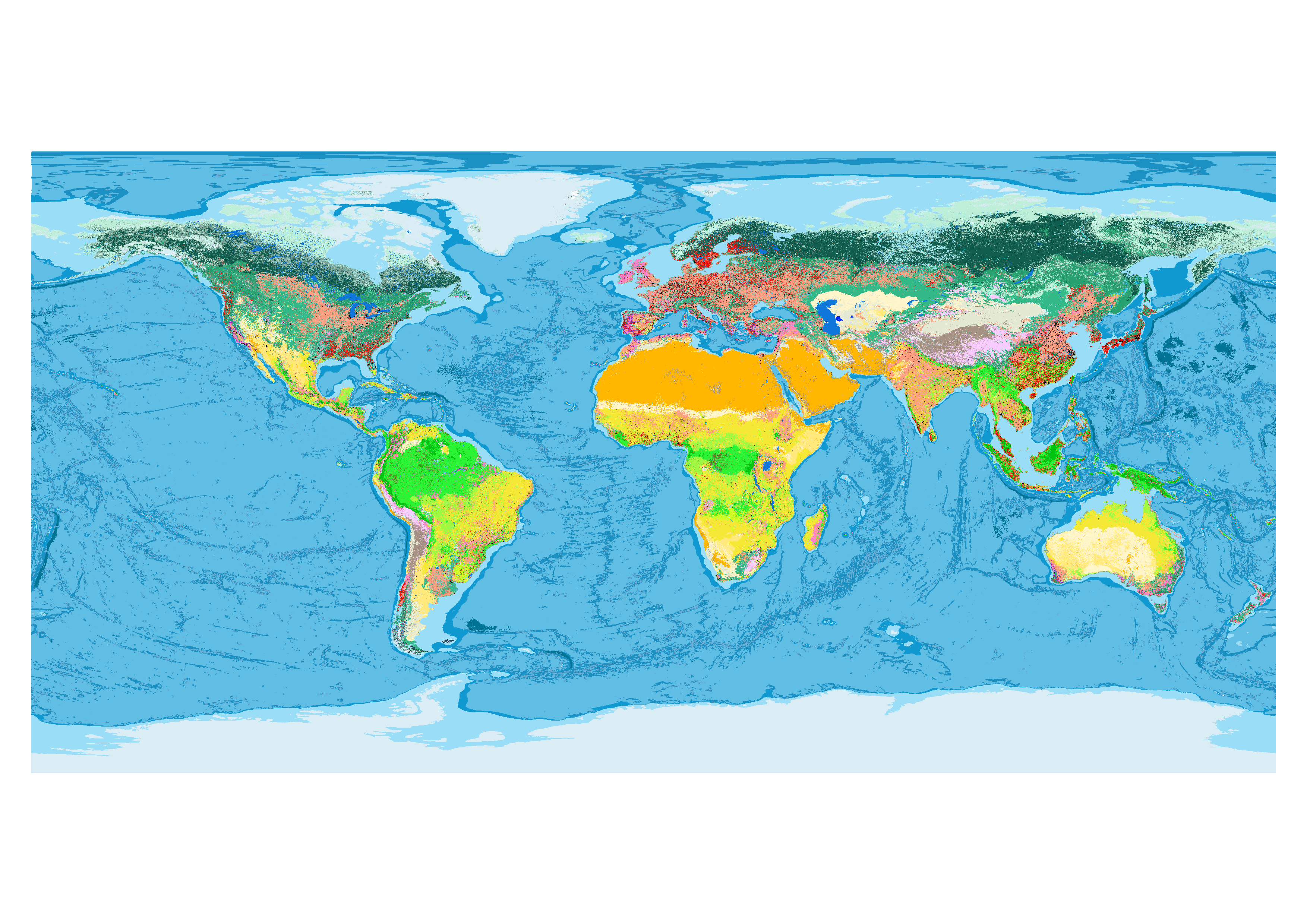
Using the Google Earth Engine platform I developed Javascript code to create the first global habitat map specifically for species habitat analyses. The map is created by intersecting best available global datasets on land cover, climate and land use. The code and map is released under a Creative Commons Attribution 4.0 International license and thus can be freely shared, used or modified. The most recent version of the habitat type layer is uploaded to Zenodo.
To run the code for creating the habitat type map an account on Google Earth Engine is necessary. No Python code was developed as part of this project. Note that while the code for the habitat type map is openly available, the underlying datasets (‘assets’ in Google Earth Engine) are not all (yet) publicly available. Assets can be shared depending on the request, however there is no guarantee that I will host all original input assets for long, since they might change as improved data becomes available.
Code | Online Viewer | Data | Manuscript
Potential distribution of land-cover classes
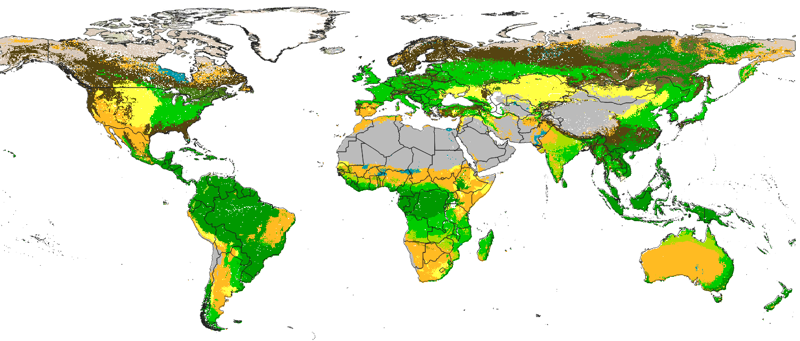
Contributed to the creation of global openly available data on the Potential distribution of land cover classes (Potential Natural Vegetation) at 250 m spatial resolution based on a compilation of data sets (Biome6000k, Geo-Wiki, LandPKS, mangroves soil database, and from various literature sources; total of about 65,000 training points). The map was created using relief and climate variables representing conditions the climate for the last 20+ years and predicted at 250 m globally using an Ensemble Machine Learning approach as implemented in the mlr package for R.
Potential global habitat mapping
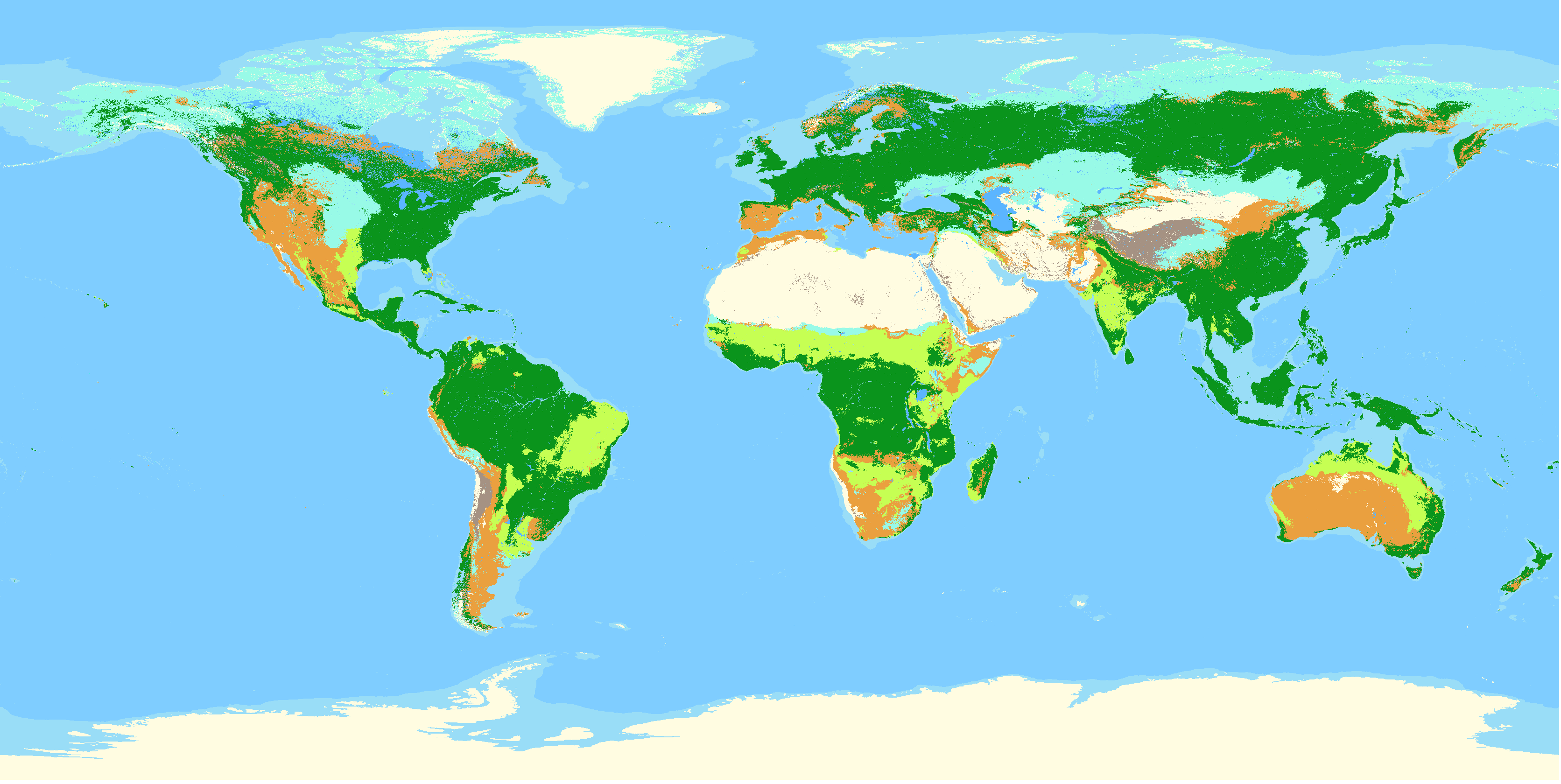
Potential global distribution, e.g. void of human influence, of habitat types following the IUCN habitat classification system at level 1. To create this layer data on the potential distribution of land cover was intersected with data on climate, elevation and topography. In total 12 classes are mapped based on the decision tree by Jung et al. (2020), with the version number of this layer matching the version of the decision tree by Jung et al. Style file for use in QGIS are supplied.
Global forest fragmentation data
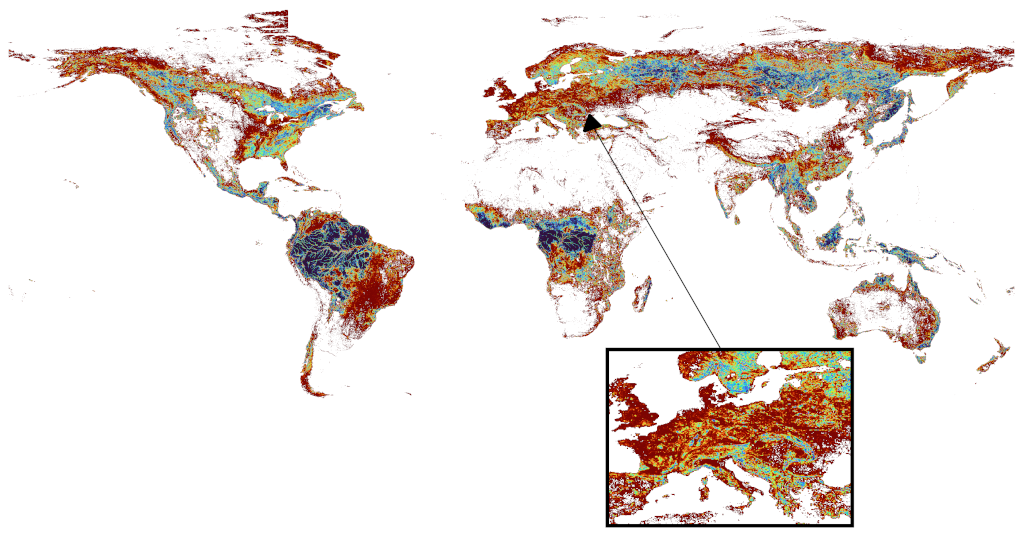
Newly created data on forest fragmentation using ALOS PALSAR data. Using the LecoS plugin I calculated on a global scale several landscape metrics at 0.1° resolution for the years 2007, 2008, 2009, 2010 and 2015. A manuscript describing the dataset was in preparation, however ultimately dropped due to changing priorities. Includes data on the mean Forest Patch Area, Patch density, Edge density, Largest patch index, Landscape division index, Fractal dimension index, Average inner edge distance, and the Percentage of Like adjacencies
Global foodscapes
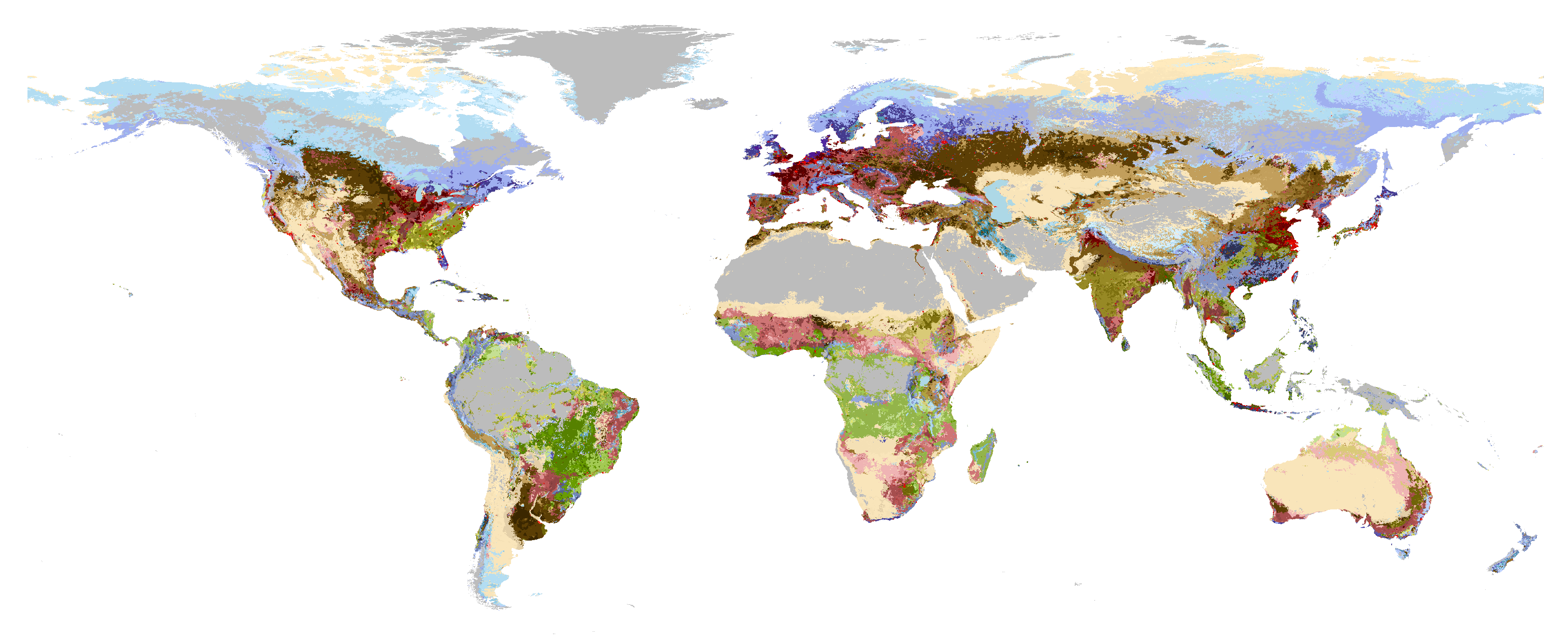 A globally consistent approximation of the world’s foodscape classes through the integration of best available global data on biophysical and socio-economic factors so as to identify a minimum set of emergent clusters of food production systems. Such a clustering of homologous systems is necessarily coarse, while still being useful to identify possible leverage points for sustainable agriculture.
A globally consistent approximation of the world’s foodscape classes through the integration of best available global data on biophysical and socio-economic factors so as to identify a minimum set of emergent clusters of food production systems. Such a clustering of homologous systems is necessarily coarse, while still being useful to identify possible leverage points for sustainable agriculture.
The data can be interactively explored and the layer is made openly downloaded here. Furthermore, as convenience, a layer further aggregated to reflect three levels of food production intensity is also provided in the same repository. A preprint describing the layer is currently in review.